Whether through symmetric or asymmetric cell division, stem cell progeny must choose between assuming a stem cell identity by remaining undifferentiated or adopting a progenitor cell identity by committing to differentiate. To successfully adopt a progenitor identity, a stem cell progeny must exit from the stem cell state and stably restrict the developmental potential. In contrast to stem cells, a progenitor cell exclusively generates differentiated cell types. Progenitors with incomplete restriction of developmental potential likely will revert into stem cell-like cells instead, and might be more susceptible to be transformed into tumor-initiating stem cells when acquired tumor driver mutations. Furthermore, ectopically activating tumor stem cells to commit to differentiate can reduce tumor burden. Thus, regulation of developmental potential lies at the intersection of normal programming of stem cell progeny in the healthy state and their abnormal re-programming in the tumorigenic state.
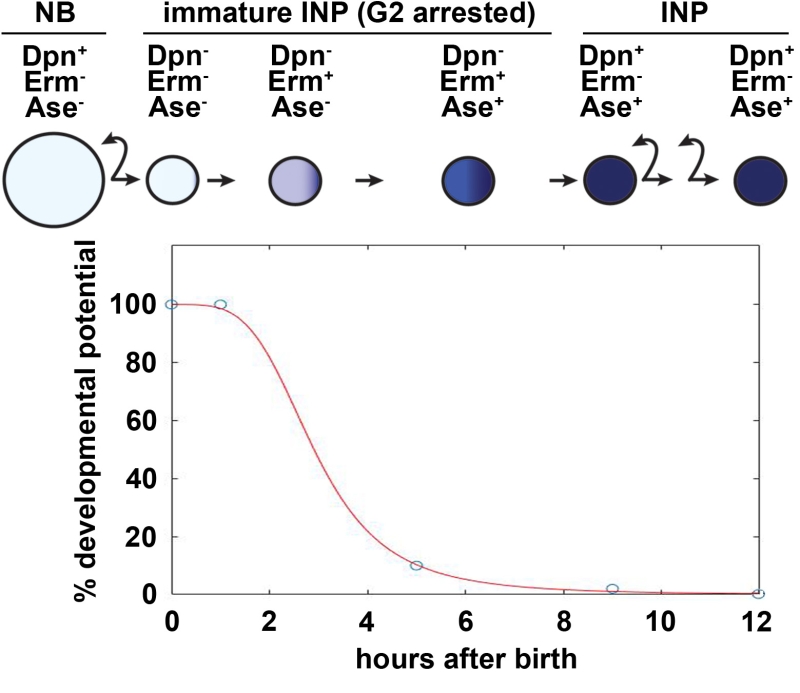
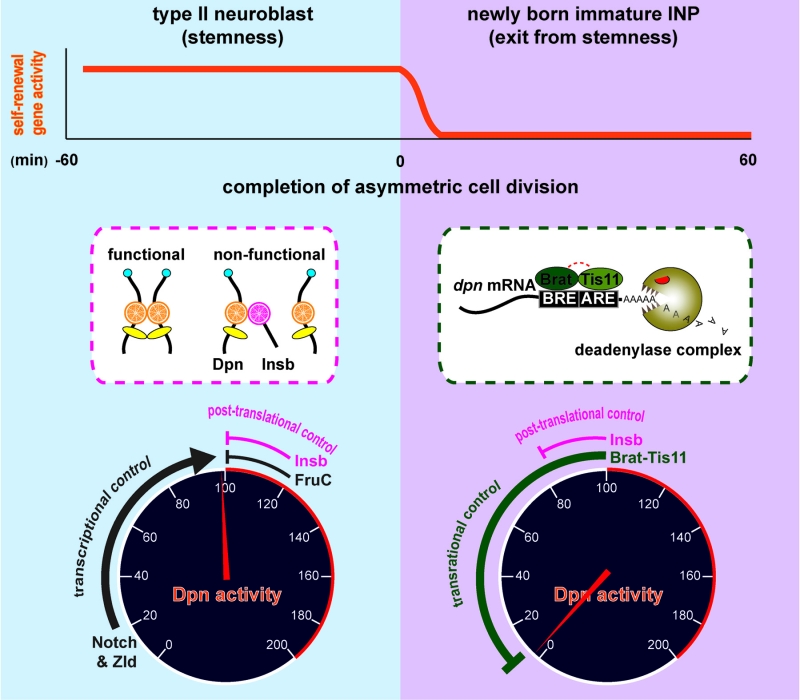
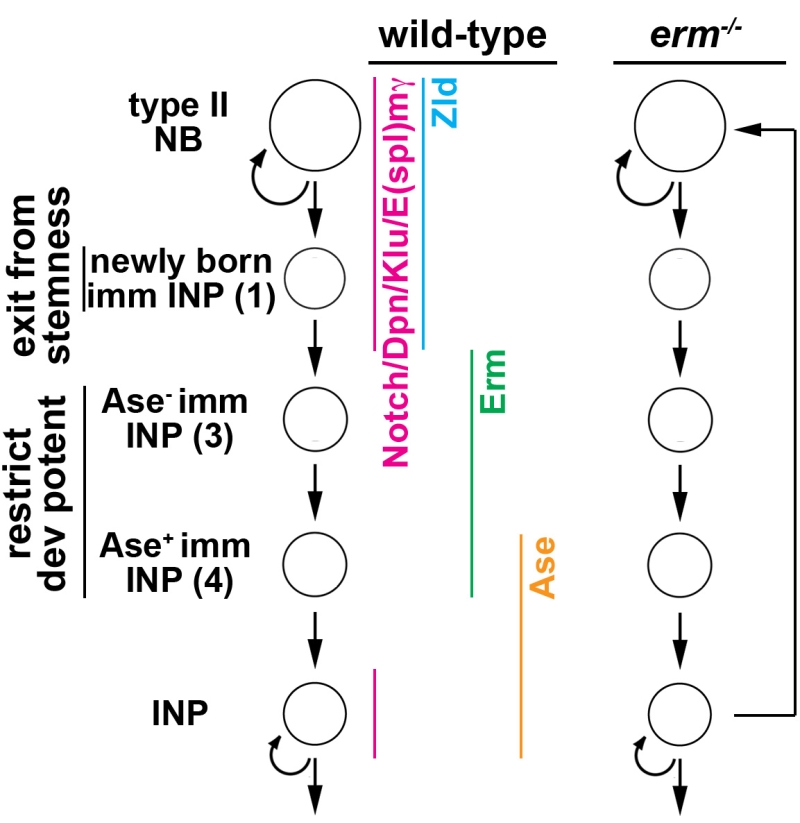
Similar to vertebrate embryonic neural stem cells, Notch signaling plays a key role in regulating the decision to self-renew or to differentiate following asymmetric type II neural stem cell (neuroblast) division in the fly larval brain (Fig. 1). The neuroblast progeny destined to commit to differentiation or the newly born immature INP exits from the neuroblast state and commits to differentiation by initiating the restriction of developmental potential less than 60 minutes after birth. 8 hours later, an immature INP adopts an INP identity and re-enters the cell cycle to exclusively generate differentiated cells. Our lab uses the type II neuroblast lineage as an in vivo paradigm to define the molecular mechanisms that
- drive timely exit from the stem cell state in the newly born immature INP
- control timely initiation of developmental potential restriction in immature INPs
- implement stable restriction of developmental potential in INPs.