Nature's chem lab: How microorganisms manufacture drugs
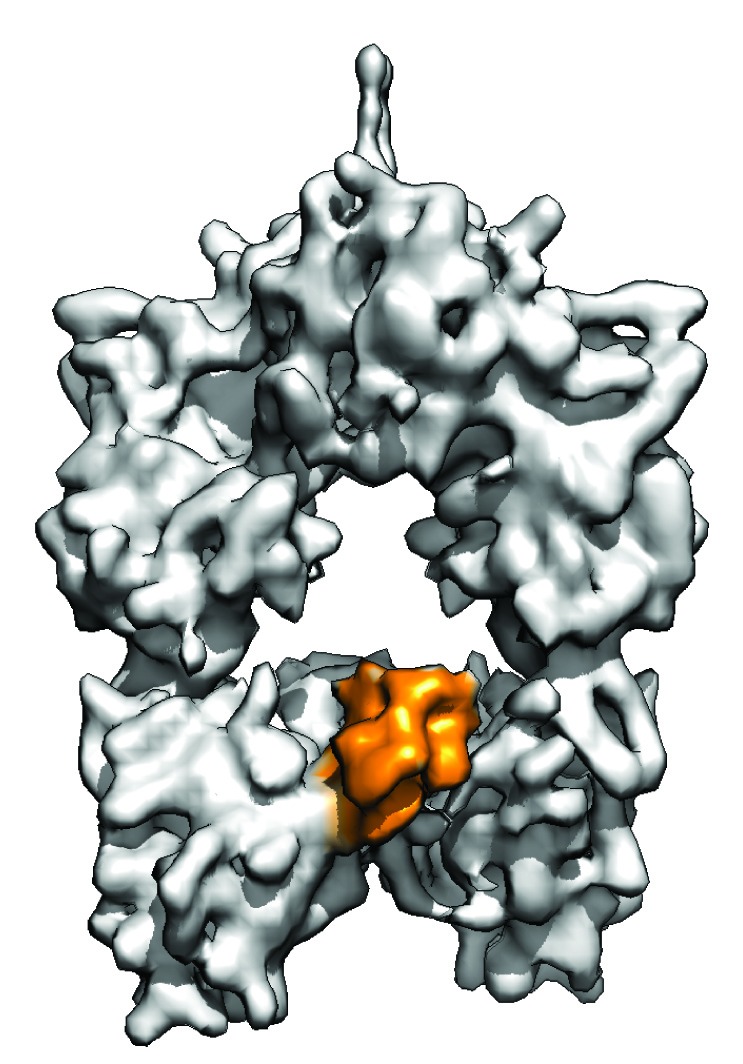
ANN ARBOR -- Researchers at the University of Michigan have obtained for the first time three-dimensional snapshots of the “assembly line” within microorganisms that naturally produces antibiotics and other drugs. Understanding the complete structure and movement within the molecular factory gives investigators a solid blueprint for redesigning the microbial assembly line to produce novel drugs of high medicinal value.
In a pair of articles published in Nature on June 18, Georgios Skiniotis, David Sherman and Janet Smith of the Life Sciences Institute (LSI) and Kristina Håkansson of the Department of Chemistry describe the structure of a central enzyme in the assembly process that creates polyketides, a broad class of diverse and bioactive chemical compounds that comprise some of the most important antibiotics, antifungal agents, cancer chemotherapeutics and immunomodulators in wide clinical use.
“Ultimately, understanding the details of polyketide creation can allow us to successfully design and engineer systems for the production of novel products with high medicinal value,” said Skiniotis, a structural biologist who is the corresponding author on both papers.
According to the Centers for Disease Control and Prevention, at least 2 million people become infected with antibiotic-resistant bacteria each year in the United States, and more than 23,000 people die as a result. Complications from infection that can’t be successfully treated with existing antibiotics lead to many more deaths.
“The threats of antibiotic resistance keep growing and we are in urgent need of new drugs,” Sherman said. “Now that we can see how the machine is working, we can understand why previous attempts to engineer these natural factories to make drugs have failed—and how we might be able to do it better.”
Nearly two-thirds of all drugs currently used are derived partially or entirely from a natural source. But the natural process used by microorganisms to create the medicines has been a “black box,” said Smith, a structural biologist who studies infectious diseases and viruses.
“We knew something went in and something else came out, but we didn’t know what happened inside,” she said. “Now we know, and having this blueprint of how microorganisms make chemicals means we can now begin to redesign the factory to make the chemicals we want for new and better drugs.”
Not what the scientists had anticipated
The polyketide synthase (PKS) enzyme was derived from a type of terrestrial bacteria called Streptomyces venezuelae, which produces pikromycin, an antibiotic precursor to the widely prescribed erythromycin. The structures published in Nature describe the enzymatic steps of the fifth cycle in a six-cycle biological assembly-line process that produces pikromycin.
“The scientific community has been waiting for decades for this structure, and the team achieved this feat with this work,” said Pieter Dorrestein of the University of California, San Diego. Dorrestein is the director of the Therapeutic Discovery Mass Spectrometry Center, co-director of the Institute of Metabolomic Medicine and a faculty member in the Skaggs School of Pharmacy and Pharmaceutical Sciences Departments of Pharmacology, Chemistry and Biochemistry. “As traditional approaches have failed to systematically develop new chemical entities, it is now time to fully understand these systems in great detail so that rational engineering may become possible; this work is a milestone toward achieving this goal.”
The structure was not what the scientists had anticipated. “What surprised us was that a tiny protein called acyl carrier protein, or ACP, moved sequentially to different sites within the PKS factory, adding additional chemical components—and each step was determined by the cargo it acquired in the previous step,” Smith said. “Understanding the role of ACP radically changes how we understand the entire structure and process.”
As an X-ray crystallographer, Smith was interested because, she said, “These machines were doing cool chemistry, but how did they do it? How did all the parts work together? I really wanted to see how.”
'A long-awaited and major breakthrough'

The work began in the lab of Sherman, a bioorganic and medicinal chemist who studies microorganisms with the goal of finding new medicines and who first started researching the synthesis of antibiotics by microorganisms in the 1990s.
“I was interested in understanding the molecular machines that make drugs,” he said. “There were lots of models proposed and lots of debate and disagreement, but very little evidence. Over 15 years we were able to characterize the basic biochemical features of this remarkable biochemical assembly line, but high-resolution features remained beyond our grasp—until now.”
The discovery published in Nature was made possible through the Skiniotis lab’s application of cryogenic electron microscopy (cryo-EM) visualization, a technology that enables researchers to obtain high-resolution snapshots of protein complexes after rapidly freezing them.
"The cryo-EM structure reported here is a long-awaited and major breakthrough towards understanding the overall organization and catalytic mechanism of these very complicated biosynthetic machines," said Barbara Gerratana of the National Institutes of Health's National Institute of General Medical Sciences, which partially funded the research. "I look forward to seeing whether these data will propel and ease the engineering of systems for the production of polyketides, which so far have not been very successful."
The cryo-EM visualizations provided the first images of the complete structure and of the changes and movement of the ACP protein that builds the chemical end product. “We had low expectations because of the size and flexibility of the enzyme,” Skiniotis said. “But when we finally figured it out and could see the ACP, it was like a switch was flipped.”
The researchers describe two years of intense collaboration in the LSI: from rushing freshly purified proteins from Smith’s structural biology lab, to bearing key chemical components from the Sherman lab, and to obtaining snapshots of the protein complexes in the cryo-EM suite a few floors down. “It helped that we were all here in the LSI and able to go in and out of each other’s labs at any given time and tap into our colleagues’ expertise,” Skiniotis said. “It’s an illustration of collaborative effort at its best.”
Final confirmation of assembly-line function and product formation came from advanced mass spectrometry experiments in the Håkansson laboratory. “It is very exciting to work as part of a big team, with four primary investigators and multiple students and postdocs from all groups involved—all with diverse, complementary expertise that enable game-changing discoveries like this,” Håkansson said.
Georgios Skiniotis is a research associate professor and the Jack E. Dixon Professor in the Life Sciences Institute, where his lab is located, and an associate professor in the Department of Biological Chemistry, U-M Medical School.
David Sherman is a research professor in the Life Sciences Institute, where his lab is located, and the Hans W. Vahlteich Professor of Medicinal Chemistry and associate dean for research and graduate education in the College of Pharmacy at U-M.
Janet Smith is a research professor in the Life Sciences Institute, where her lab is located, and the director of the Center for Structural Biology, housed at LSI, and the Margaret J. Hunter Collegiate Professor of Life Sciences, Department of Biological Chemistry, U-M Medical School.
Kristina Håkansson is a professor in Department of Chemistry in the College of Literature, Science, and the Arts.
Go to Articles
- Structure of a modular polyketide synthase, Nature. DOI: 10.1038/nature13423
- Structural rearrangements of a polyketide synthase module during its catalytic cycle, Nature. DOI: 10.1038/nature13409


